 
It uses the P BAD promoter, a tightly regulated bacterial promoter system, which gives minimal expression in the absence of inducer and toxicity by misfolded protein is readily controlled. In the current method, we prefer the araBAD expression system for production of membrane proteins. There are wide varieties of plasmids, which can be used for protein expression in E. Ho, Bert Poolman, in Methods in Enzymology, 2015 3.1 Construction of expression vector for target protein–GFP–ErmC fusion 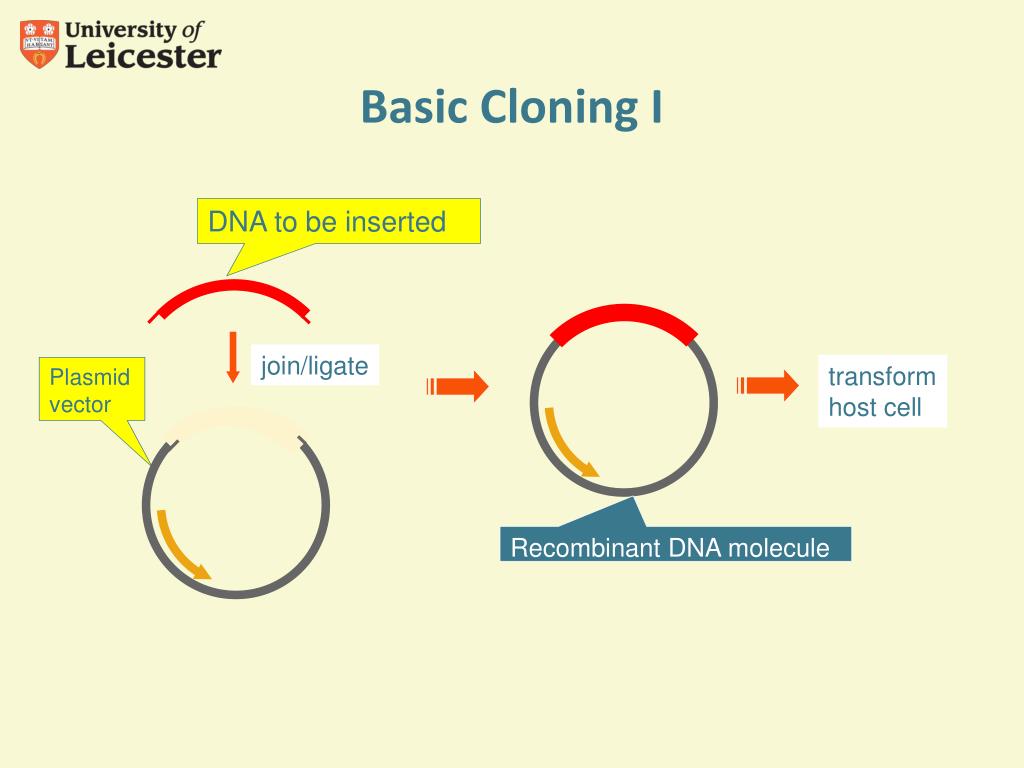
The constructs can be validated by sequence analysis at this stage or the analysis can be performed after analysis for expression and solubility.įranz Y. The preparation of LIC compatible vector is similar to the above procedure but uses the complementary base dGTP ( Eschenfeldt et al., 2009). coli cells and select for transformed colonies ( Sambrook and Russell, 2001).įollowing the heating process the LIC plates are stored in the refrigerator at 4 ☌ until needed.  
Use the entire annealing reaction to transform ~ 50 μL competent E. Incubate the annealing reaction 5–10 min at room temperature. In another 96-well plate, anneal 1–2 μL of the T4-Polymerase-treated PCR LIC fragments with 4 μL (20–50 ng) T4-Polymerase-treated LIC vector. Incubate on a heat block at 75 ☌ for 20 min to inactivate the T4 DNA polymerase. Our studies of various fragment-to-vector ratios ( Dieckman et al., 2006) indicate a wide tolerance for variation in the amount of target DNA fragment on the annealing reaction. 5Īdd 30 μL (40–100 ng) of purified PCR fragment to the LIC reaction mix and pipette up and down several times to mix. 4Īrray 10.4 μL of the LIC reaction mix into a polypropylene 96-well plate. Pipette up and down several times to uniformly distribute the enzyme in the reaction mix. Keep this mixture on ice and add the T4 DNA polymerase just before use. dĢ50 units of T4 DNA Polymerase (LIC quality, ~ 2.5 units/μL, EMD Biosciences/Novagen). cĢ28 μL of 100 m M dithiothreitol (DTT) solution (Novagen cat. bĤ65 μL of 25 m M dCTP, molecular biology grade (Promega cat. Our comparison of various common T4 DNA polymerase reaction buffers shows less than a 25% difference in the cloning efficiency of the final product. T4 DNA polymerase reaction buffers supplied by most vendors can be substituted for the LIC reaction buffer described in this section. Make LIC Reaction mix sufficient for one 96-well plate by combining the following reagents: aĤ65 μL of 10 × T4 DNA polymerase buffer. The specified method is scaled to microwell plates (96 targets) but can be adjusted to any number of targets. LIC versions of a range of transfer plasmids are now also available (i.e., pBAC-2cp Ek/LIC, Novagen), for rapid, directional cloning of PCR products.Īppend the appropriate LIC specific nucleotide sequences (e.g., forward primer: TACTTCCAATCCAATGCC, reverse primer with added stop: TTATCCACTTCCAATGTTA) to the target specific PCR primers. The noncovalently joined molecules are then repaired in E. coli to produce a seamless fusion. When the vector is combined with the insert, the poxvirus DNA polymerase 3′–5′exonuclease activity converts the double-stranded extensions into short single-stranded sequences and fuses these regions to the corresponding ends of the linearized vector. The 3′ and 5′ regions of homology are generated by adding 15 bp extensions to both PCR primers that precisely match the ends of the linearized vector. The foreign gene is amplified by PCR and its single-stranded ends are fused to the homologous ends of a linearized vector using a poxvirus enzyme. The In-Fusion™ PCR Cloning System (BD Biosciences) is based on LIC and utilizes several transfer plasmids based on pBacPAK8 with C- and N-terminal 6× His tags. Ligation-independent cloning (LIC) is a method that utilizes short homology arms for the directional cloning of PCR products negating the need for restriction enzyme digestion or ligation reactions. King, in Comprehensive Biotechnology (Third Edition), 2011 1.22.10.1.1 Ligation-Independent Cloning 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
